In this page you will find information about the predictor implemented in ProABC-2.
Convolutional Neural Network (CNN)
The neural network used by proABC-2 consists of three convolutional modules (Conv11, Conv12 and Conv2), a fully connected feed-forward module (Ff1) and an output layer. Conv11 and Conv12 are identical and consist of three parts; a 1D convolutional layer with 32 filters of size 3x1 and a stride of 1, followed by a 1D max pooling layer of size 10x1 and a stride of 3. Conv2 also consists of three parts; a 1D convolutional layer with 64 filters of size 3x1 and a stride of 1, followed by a 1D average pooling layer of size 6x1 and a stride of 3. Ff1 consists of a fully connected layer with 512 nodes. The final output layer has for each of the 297 residues 3 nodes, predicting the general interactions, H-bonds and hydrophobic interactions, amounting to 891 nodes.
These modules are combined in the following way. The one-hot encoded heavy and light chains are connected to Conv11 and Conv12 respectively. The extracted features of the heavy and light chains are then concatenated and enter Conv2 for a deeper feature extraction. The final extracted features from Conv2 are then flattened (reduced to one dimension) and concatenated with the additional features (germline, loop lengths and canonical structures) before entering Ff1 and finally into the output nodes. The purpose of Ff1 is to learn each individual residue’s role in the paratope based on the extracted features and the additional ones. The architecture is shown in the figure below. Exponential Linear Units (ELU) were used as activation functions for Conv11, Conv12, Conv2 and Ff1, and sigmoid on the final output.
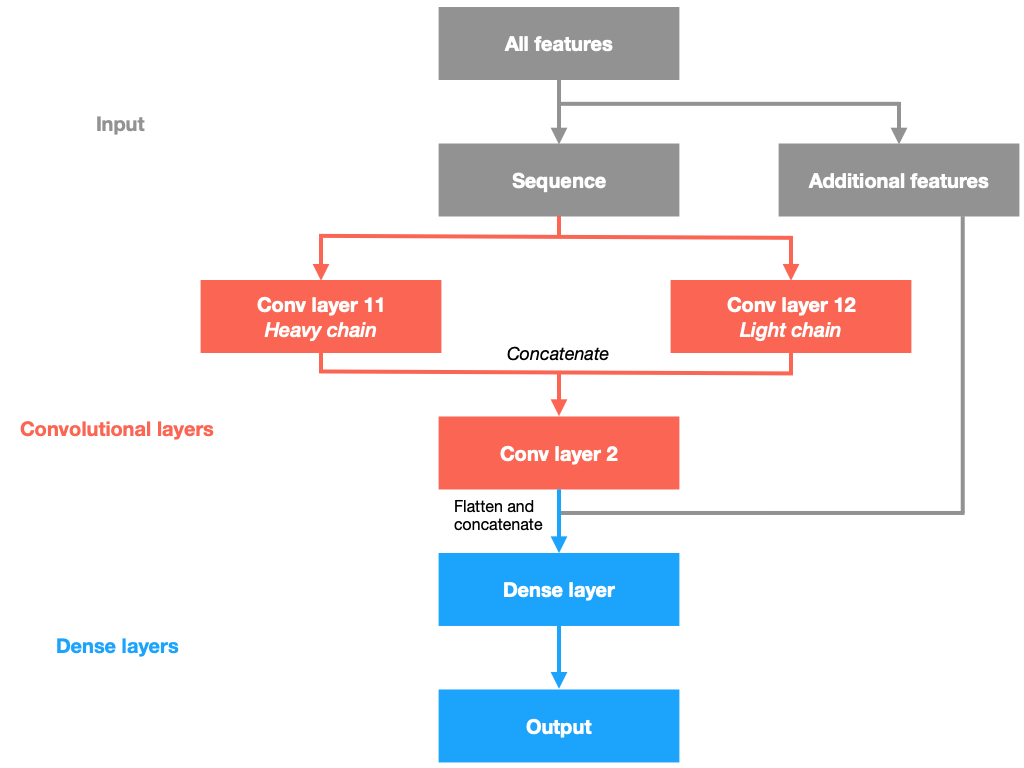
Performance
The evaluation of the model has been performed using 10-fold nested cross validation on the full training set of antibody-antigen complexes (769 complexes). The performance was measured taking into account three different metrics: area under the Receiver Operating Characteristic curve (AUC), Matthew Correlation Coefficient (MCC) and F-score. MCC and F-score were calculated using a threshold of 0.40, 0.30 and 0.30, respectively for all the general interactions of the paratope (Pt), hydrophobic interactions (Hy) and for hydrogen bonds (Hb). These cut-offs were selected by averaging the thresholds that for each fold of the cross validation gave the best MCC.
In the table below si summarised the proABC-2 performance for the three different types of interaction.
| Interaction | AUC | MCC | F-score |
|---|---|---|---|
| Pt | 0.96 | 0.57 | 0.59 |
| Hy | 0.95 | 0.44 | 0.41 |
| Hb | 0.94 | 0.33 | 0.37 |





